Nathan Sheffield, PhD

Lab Page: http://www.databio.org
I use computational and experimental tools to study high-throughput cancer epigenomic data. My biological focus is on Ewing sarcoma, a childhood cancer with metastatic cases having low long-term survival.
Ewing sarcoma is almost always driven by a single, well-characterized mutagenic event: usually a chromosomal translocation leading to the fusion protein EWS-FLI1. EWS-FLI1 is a potent driver oncoprotein, and it completely rewires a cell’s regulatory state by activating Ewing sarcoma specific enhancers and indirectly repressing other developmental regulators, pushing the cells from a mesenchymal state into a more stem-cell-like diseased state.
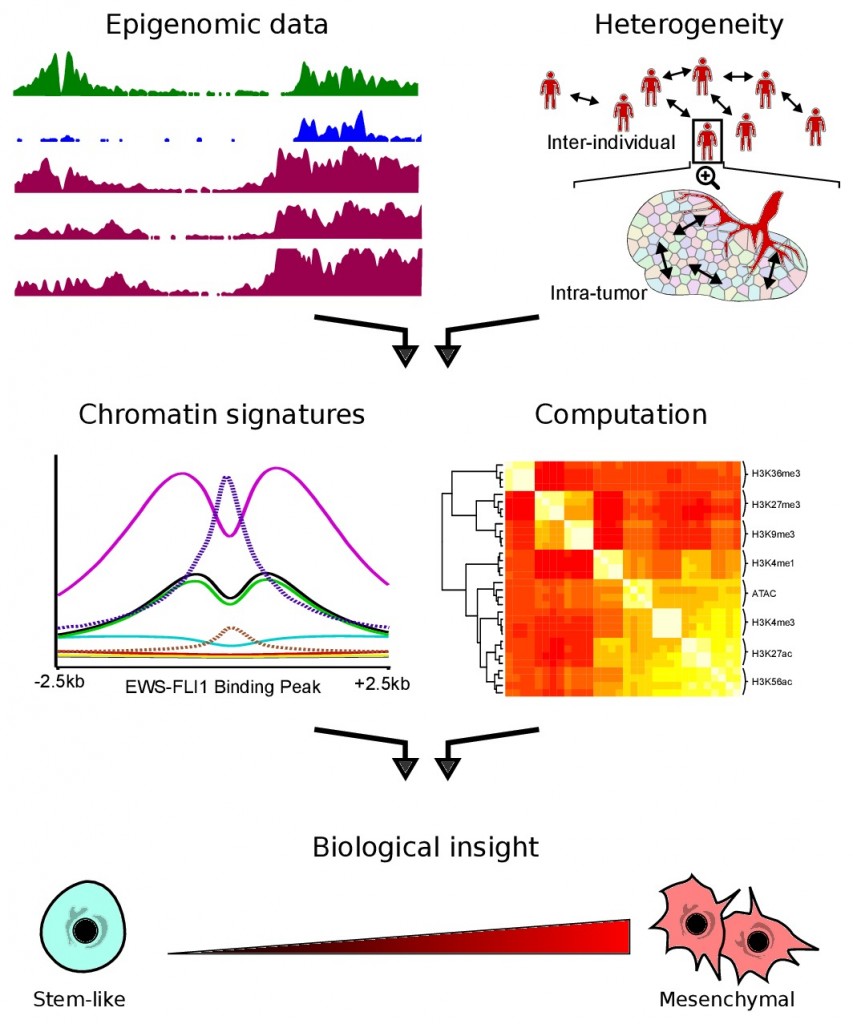
I study epigenomic signatures (like DNA methylation, histone modification, and chromatin organization) of Ewing sarcoma to better understand how EWS-FLI1 causes cancer. What regulatory systems and pathways are disrupted by the EWS-FLI1 fusion? How do within-tumor and between-individual heterogeneity affect this disruption? I collect genome-scale data in single cells and cell populations, and then harness the power of supercomputing, machine learning, and software engineering to answer these types of biological questions. This research increases our understanding of the basic biological processes of differentiation and carcinogenesis, and will ultimately lead to new ways to target and combat the disease.