Research Cores
The PCEN supports two biomedical research core laboratories to assist investigators in two important areas: 1) Single-cell transcriptome and genome analyses and 2) Genome Mapping, Manipulation, Analysis, and Bioinformatics Core.. The core laboratories are directed by experienced researchers and staffed with highly trained personnel.
The services provided by these cores enable PCEN investigators to carry out their research in a cost-effective and efficient way. Each core unit serves as a resource for all PCEN two main research projects and pilot projects at all times. In addition, these laboratories serve other investigators within the School of Medicine and the Department of Biology at UVA, as well as researchers from other institutions. Each core offers a variety of specialized and innovative research services, and provide hands-on training to scientists as well as consultation services and access to state-of-the-art research equipment for experienced users.
The scope of these cores is in line with the overall theme of this PCEN: “Kidney Development and Disease: Cell Fate and Precursors of Disease in the Young and Adult.” The experimental approaches of the PCEN involve examination of the mechanisms that control cell fate, plasticity, and identity and the cellular signals underlying the phenotypic transformation that occurs during kidney development, injury and repair. The cores provide specialized technologies and expertise needed to accomplish the goats of the PCEN.
Director: Mazhar Adli, PHD
Most of the projects within the PCEN require Epigenomic and Genomic tools and methodologies, including epigenomic and genomic editing, Assay for Transposase Active Chromatin followed by sequencing (ATAC-seq), application of CRISPR-based approaches to modify the genome or re-write the epigenome of renal cells in a locus- and gene-specific-specific manner. The major focus of the core is on epigenomics and editing methodologies which are suitable for those investigators within and outside the center by devoting energy and resources to a combination of methodologies that capitalize on the outstanding expertise on epigenetics and epigenetic re-writing combined with specific bioinformatics tools designed to deal with the fields of epigenomics and kidney development. The main Aim of this core is to provide technical and analysis expertise in areas of epigenome mapping, locus specific genome and epigenome editing and genome-scale data analysis.
Dr. Mazhar Adli serves as the Core Director. The proposed core activities are based on Dr. Adli’s extensive and unique expertise. During his postdoctoral training at Harvard Medical School and the Broad Institute, Dr. Adli established a unique technical and analytical expertise in genomics and epigenomic profiling and computational data analysis. During this process, he developed the Nano-ChIP-Seq technology that overcame the cell number limitation of conventional ChIP-Seq technology. In addition, he played critical roles in multiple large-scale projects including Roadmap Epigenome Mapping Consortium and cancer genome projects.
In his own laboratory at the UVA Department of Biochemistry and Molecular Genetics, Dr. Adli is combining genome and epigenome mapping expertise with novel CRISPR-based manipulation tools to achieve his lab’s ultimate research goals. To this end, his laboratory has developed a strong expertise in CRISPR technologies. The Adli lab is actively developing and utilizing novel CRISPR tools to edit genome, manipulate epigenome and image live cells chromatin dynamics in living cells.
Through the core, Dr. Adli shares the technical as well as the analytical expertise of his laboratory with all members of the PCEN to provide support in experimental design, techniques and protocols, and subsequent analytical pipelines for these cutting edge tools.
Director: Silvia Medrano PhD
It is becoming clear that a high level of cell-to-cell transcriptome and proteome variation exists in cell populations from in vivo and in vitro sources. For example, during differentiation, each cell makes fate decisions independently by receiving and processing signals from other cells and executing complex gene regulatory programs. As a result, studies conducted in bulk can overlook important cellular heterogeneity and lead to false conclusions. Single-cell approaches avoid this problem by allowing the dissection of heterogeneous cell populations in complex organs like the kidney. Therefore, single-cell resolution of transcript and protein expression dynamics is crucial to understand the phenotypic changes that occur during kidney development, injury and repair.
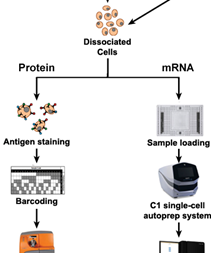
Transcript-level dynamics in individual cells can be measured by scRNA-Seq using the Fluidigm C1 System. In addition, the Fluidigm CyTOF 2 single-cell mass cytometry system is a powerful proteomics platform that enables the simultaneous analysis of over 30 proteins in individual cells.
The work to be conducted by the SCGMC core is divided in two Aims.
Aim 1: To provide support and services for genome, transcriptome and functional/ phenotypic analysis at thesingle-cell level. The core offers high quality, cost effective mRNA-sequencing, DNA-sequencing, qPCR pre-amplification of genes panels, and mass cytometry services for all PCEN investigators and researchers in other institutions. These services include performing specialized single-cell research techniques, consulting in experimental design, supporting preparation of manuscripts and grant proposals, training, and accessing state-of-the-art equipment.
Aim 2: To apply the knowledge and expertise of the core to explore new areas with translational potential. The core supports clinical and translational research by performing single-cell analysis of samples from patients, i.e. cells from urine, blood and biopsies, with the ultimate goal of developing novel diagnostics and therapeutic interventions for kidney diseases in children.
SCGMC core services are used by all PCEN investigators as their individual projects heavily rely on state-of-the-art single-cell strategies. In addition, the core provide services to investigators holding awards from the Center’s Pilot and Feasibility Program and independently funded users. Integration of both mRNA and protein expression analysis, at the single-cell level, provides much greater resolution of the dynamics of cell fate decisions during kidney development and disease.